Prof. Dr. Herre Jelger Risselada
Department of Physics
Otto-Hahn-Str. 4
44227 Dortmund
E-Mail: jelger.risselada@tu-dortmund.de

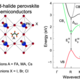
"The research group employs molecular simulation techniques to tackle relevant problems in the field of biophysics and related molecular fields within a multidisciplinary and interdisciplinary research environment (e.g., the Resolv excellence cluster). Particularly, we focus on the development of enhanced sampling and multiscale methods to study phenomena occurring at fluid (e.g., biological lipid membranes) and solid interfaces (e.g., metal interfaces and graphene). An important current focuses is on a subject coined 'facial recognition of fluid interfaces', i.e., how can (bio)molecules optimally recognize and bind fluid interfaces despite their highly disordered and dynamic nature. To this aim, we develop evolutionary molecular dynamics simulations (evo-MD) methods which couple evolutionary algorithms (sampling of chemical space) to coarse-grained molecular dynamic simulations (sampling of phase-space) to resolve the global optima in chemical space. In essence, evo-MD enables molecules to adapt themselves to optimally recognize fluid interfaces in the course of a simulated evolution. Our research may yield unique insights on how biomolecules such as peptides and proteins recognize distinct fluid features in biological lipid membranes with interesting applications for the fields of drug design, surfactant design, and the design of (bio)sensors."